

 technology
technologyThe Biomedical Knowledge Miner (BiK>Mi) provides tools to access and validate knowledge encompassing all of the latest information pertaining to COVID-19. Our goal is to integrate as much relevant information as we can into a single, comprehensive model whereby users are able to investigate all-known molecular interactions pertaining to SARS-CoV-2 as well as possible therapeutic routes.
The department of Bioinformatics at the Fraunhofer Institute for Algorithms and Scientific Computing (SCAI) seeks to reshape scientific knowledge into a single, computable repository despite the fact that this knowledge is stored in a variety of formats, databases, and sources. To ensure that our knowledge graph contains the latest findings, chosen publications were manually curated and their contents converted in Biological Expression Language (BEL) statements. These statements are then parsed and used to build an interactive knowledge graph by the in-house developed software package e(BE:L). Information provided here includes also includes publicly available data that was reformatted and added to our model.
This project aims to construct a comprehensive computational model of biological mechanisms in the context of COVID-19 in order to more efficiently identify potential therapeutic targets and affected pathways.
Biological Expression Language (BEL) is a domain-specific language that can be used to describe biological relationships in a computable format while still maintaining the contextual information. The interactions represented within our knowledge graphs were manually converted into BEL by our team of curators. Below is an example of a BEL encoding and how the resulting BEL statement is structured. More information about BEL can be found at the BEL.bio website.
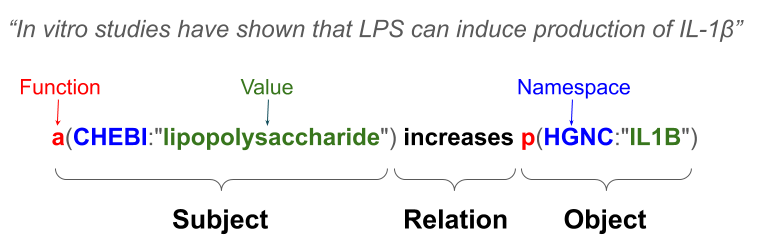
BiK>Mi and e(BE:L) are under continuous development by the Fraunhofer Institute for Algorithms and Scientific Computing (SCAI), Department of Bioinformatics, and built in the context of the PHAGO project funded by the IMI.
In addition to their AETIONOMY and PHAGO projects, IMI is also contributing to the fight against COVID-19 (read more here).